polars-bio 0.31.0: VCF Zarr Support for Array-Native Variant Analytics
polars-bio 0.31.0 adds first-class support for VCF Zarr, giving Python users a fast, lazy, DataFusion-backed way to query VCF-derived Zarr stores with Polars.
polars-bio 0.31.0 adds first-class support for VCF Zarr, giving Python users a fast, lazy, DataFusion-backed way to query VCF-derived Zarr stores with Polars.
Genomic analyses in Python typically start with reading BAM, VCF, or FASTQ files into memory. The choice of library for this step can have a dramatic impact on both wall-clock time and memory consumption — especially as datasets grow to tens or hundreds of millions of records.
pysam has long been the go-to Python library for working with these formats. It provides comprehensive bindings to htslib and is battle-tested across thousands of projects. However, several newer libraries have emerged that leverage Apache Arrow columnar format and Rust-based parsers to offer potentially better performance.
In this post, we benchmark four Python libraries head-to-head on real-world genomic data to find out which offers the best combination of speed and memory efficiency for reading BAM, VCF, and FASTQ files.
We're excited to announce significant performance improvements to GFF file reading in polars-bio 0.15.0. This release introduces two major optimizations that dramatically improve both speed and memory efficiency when working with GFF files:
Projection Pushdown: Only the columns you need are read from disk, reducing I/O overhead and memory usage. This is particularly beneficial when working with wide GFF files that contain many optional attributes.
Predicate Pushdown: Row filtering is applied during the file reading process, eliminating the need to load irrelevant data into memory. This allows for lightning-fast queries on large GFF datasets.
Fully Streamed Parallel Reads: BGZF-compressed files can now be read in parallel with true streaming, enabling out-of-core processing of massive genomic datasets without memory constraints.
To evaluate these improvements, we conducted comprehensive benchmarks comparing three popular data processing libraries:
All benchmarks were performed on a large GFF file (~7.7 million records, file and index needed for parallel reading) with both full scan and filtered query scenarios to demonstrate real-world performance gains.
For pandas and polars reading, we used the following methods (thanks to @urineri for the Polars code).
Since Polars decompresses compressed CSV/TSV files completely in memory as highlighted here, we also used polars_streaming_csv_decompression, a great plugin developed by @ghuls to enable streaming decompression in Polars.
Test query used for filtered benchmarks (Polars and polars-bio):
result = (
lf.filter(
(pl.col("seqid") == "chrY")
& (pl.col("start") < 500000)
& (pl.col("end") > 510000)
)
.select(["seqid", "start", "end", "type"])
.collect()
)
The above query is very selective and returns only two rows from the entire dataset.
Complete benchmark code and results are available in the polars-bio repository.
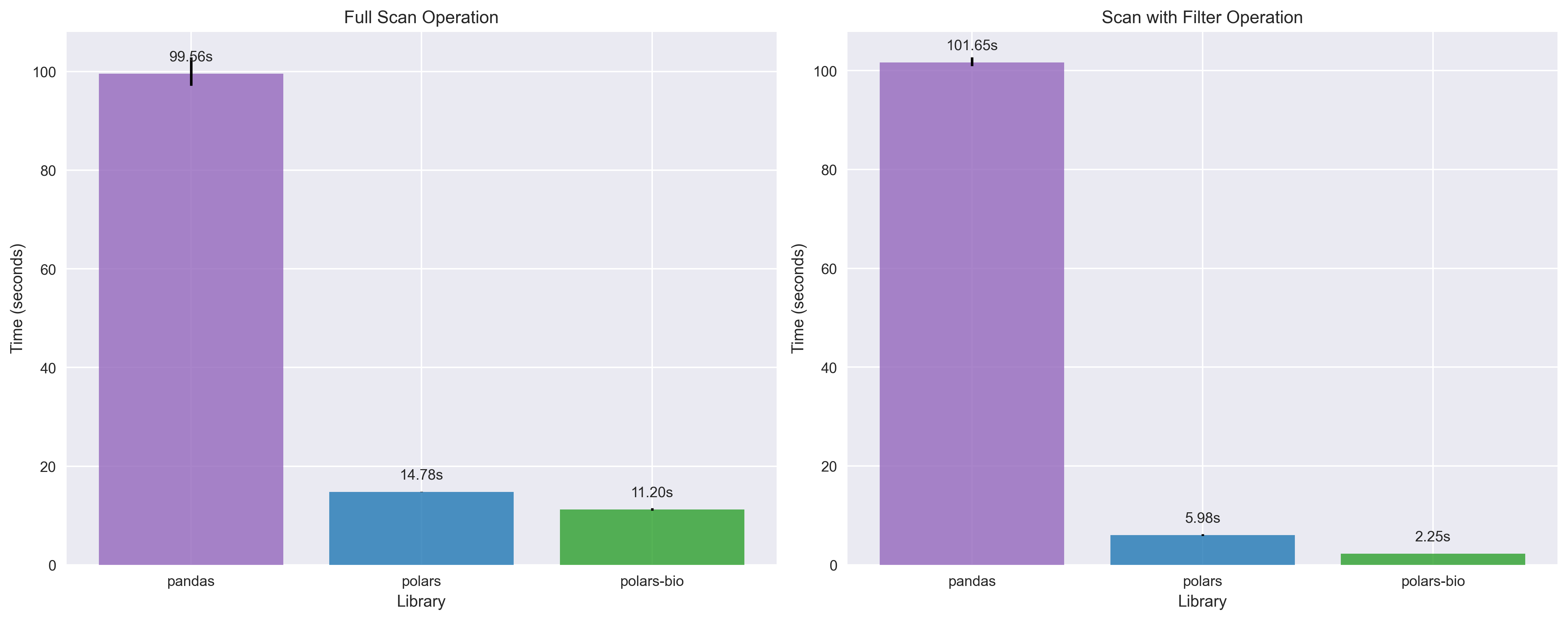
Key takeaways:
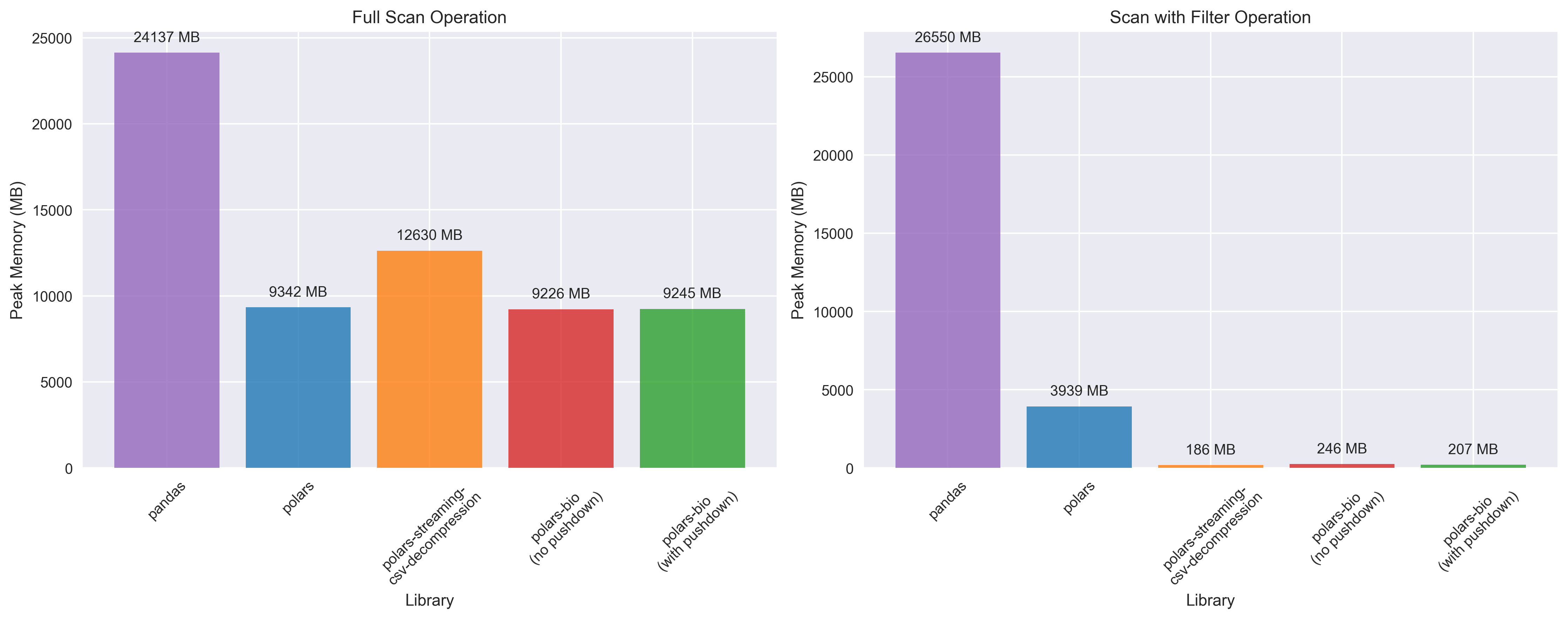
Key takeaways:
polars_streaming_csv_decompression can use more than 20x less memory than vanilla Polars and more than two orders of magnitude less memory than Pandas for operations involving filtering.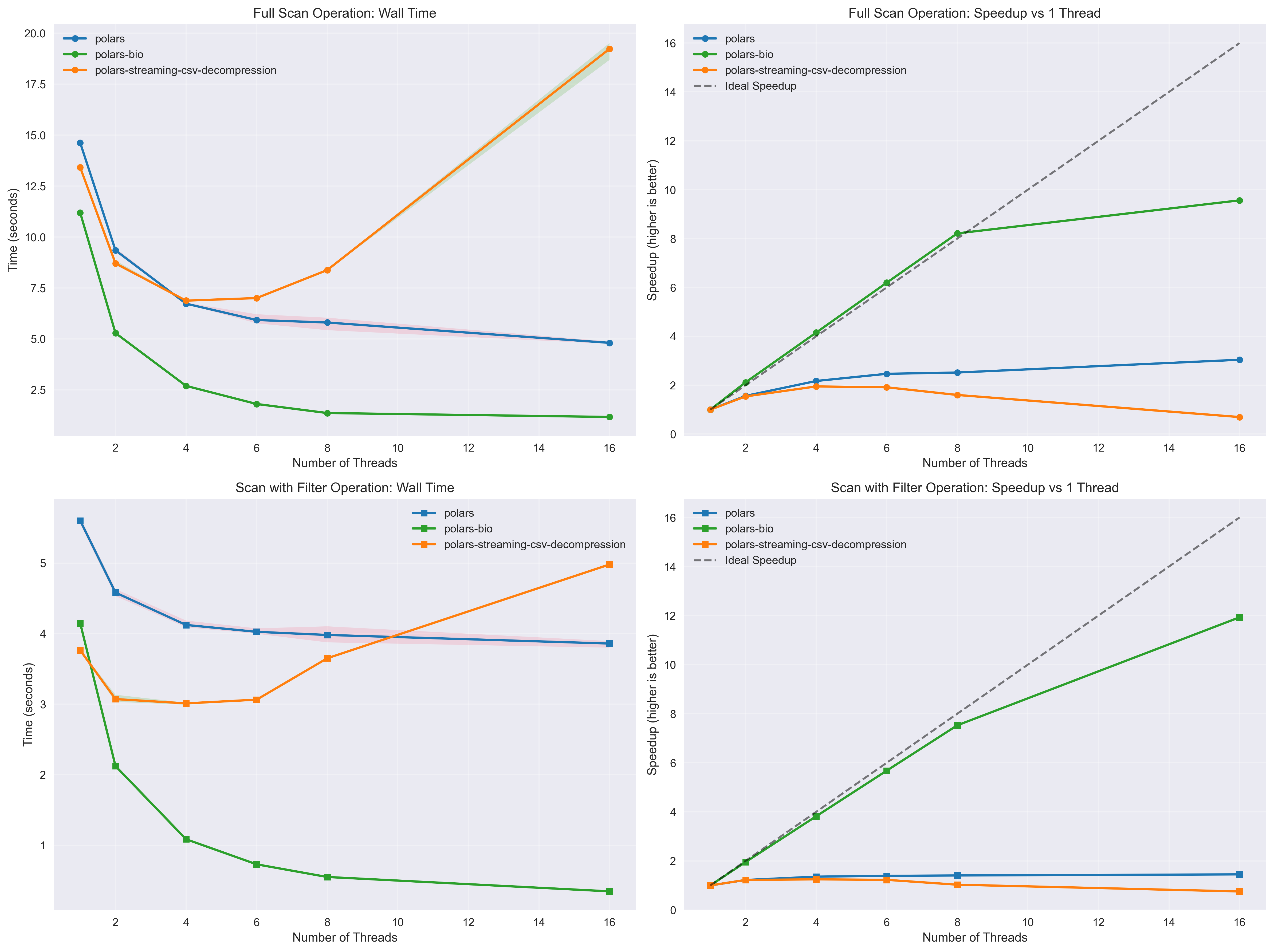
Key takeaways:
The benchmarks demonstrate that polars-bio 0.15.0 delivers significant performance improvements for GFF file processing. These optimizations, combined with near-linear thread scaling and fully streamed parallel reads, make polars-bio an ideal choice for high-performance genomic data analysis workflows.
If you haven't tried polars-bio yet, now is a great time to explore its capabilities for efficient genomic data processing with Python! Join our upcoming seminar on September 15, 2025, to learn more about polars-bio and its applications in genomics.
